Endogenous viral elements (EVEs), known as molecular fossils of ancient viral infections, are viral genetic fragments integrated into host genomes and transmissible across generations, providing a unique perspective for studying the evolutionary arms race between viruses and their hosts.
A research team led by Prof. LI Fuhua from the Institute of Oceanology of the Chinese Academy of Sciences (IOCAS), alongside collaborators from Thailand National Center for Genetic Engineering and Biotechnology (BIOTEC) and National Science and Technology Development Agency (NSTDA), carried out a series of studies, providing new insights into dynamic viral endogenization in crustaceans.
The latest study has unraveled the pervasive and dynamic integration of EVEs in crustacean genomes, including those from IHHNV and DIV1, revealing critical insights into the long-term co-evolutionary arms race between crustaceans and their viral pathogens, and highlighting urgent challenges for aquatic disease detection.
The researchers analyzed 105 crustacean genomes and identified 252 IHHNV-derived EVEs, split between ancient genomic integrations and rare modern insertions. These elements exhibit strict host specificity, found only in Decapoda, Thoracica, Isopoda and Copepoda, with Decapoda showing the highest prevalence (82.9%) and a marked expansion of IHHNV-EVEs, reflecting an intensified evolutionary struggle with the pathogen. The viral non-structural protein 1 (NS1) accounts for 76.6% of all IHHNV-EVE integrations, linking element formation closely to viral replication processes. Ancient IHHNV-EVEs were found to have ancestral inheritance traits, with collinear elements in giant isopods dating back ~245 million years and those in Penaeidae predating 100 million years, while three ancient EVEs are stably integrated in Litopenaeus vannamei across all wild and cultured populations. Only one authentic recent IHHNV integration was confirmed in Penaeus monodon, proving such insertions are ongoing but sporadic in crustaceans. Expression analyses revealed IHHNV-EVEs are weakly expressed across L. vannamei tissues, with higher levels in gills, gonads and the Oka lymphoid organ—key targets of IHHNV infection—hinting at potential vertical inheritance.
They also uncovered a critical caveat for crustacean disease detection: a high proportion of healthy L. vannamei (66.0%) and Macrobrachium rosenbergii (98.5%) tested positive for DIV1 via the WOAH-recommended ATPase-PCR assay, despite showing no pathognomonic DIV1 lesions or disease signs. While the PCR amplicons matched the pathogenic DIV1 sequence, bioassays with inoculums from these PCR-positive healthy decapods caused no mortality, viral replication or pathological damage, unlike inoculums from diseased individuals. Transmission electron microscopy confirmed no intact DIV1 virions in healthy specimens, and metagenomic sequencing revealed only fragmented DIV1 sequences (602 mappable reads out of ~27 million) covering just 33% of the viral genome—with no complete genomic sequence detected. In situ hybridization further distinguished these PCR-positive healthy samples, with signals confined to nuclei of gill and hepatopancreas cells, unlike the cytoplasmic signals in target tissues seen in genuine DIV1 infections. These false-positive DIV1 diagnoses might be caused by DIV1 viral inserts in host genomes as EVEs.
Complementary genomic analyses have expanded these findings, establishing a standardized bioinformatics pipeline for EVE identification and classification in crustaceans, and revealing universal yet lineage-specific viral endogenization across 105 crustacean species. EVE integration is found to be a ubiquitous process in crustaceans, but with distinct patterns across taxa: Decapoda primarily harbors EVEs from Nimaviridae and Retroviridae, Isopoda and Amphipoda from Adintoviridae, and Thoracica from a diverse range of viral families including Nimaviridae, Parvoviridae and Iridoviridae. Notably, all EVE integrations strictly align with known host-pathogen relationships, with no cross-species viral insertions detected, and EVE flanking regions show lineage-specific enrichment of simple sequence repeats (SSRs) and transposable elements (TEs), proving EVE integration is not random, but a genomic imprint of adaptive host-pathogen co-evolution.
Collectively, these studies redefine viral genomic colonization in crustaceans as a continuous, lineage-specific process, where ancient EVE fixation coexists with ongoing modern integrations. EVEs emerge not only as molecular fossils of past host-pathogen conflicts but also as active players in contemporary evolutionary battles, with some conferring adaptive antiviral resistance. The findings also highlight critical practical implications: EVE-associated false PCR positives threaten aquaculture trade, demanding revised disease detection protocols that combine genetic testing with histological and functional validation. For shrimp aquaculture, these insights pave the way for EVE-based biomarkers and novel antiviral strategies, while deepening our understanding of the dynamic genomic co-evolution that shapes crustacean-virus interactions across millions of years.
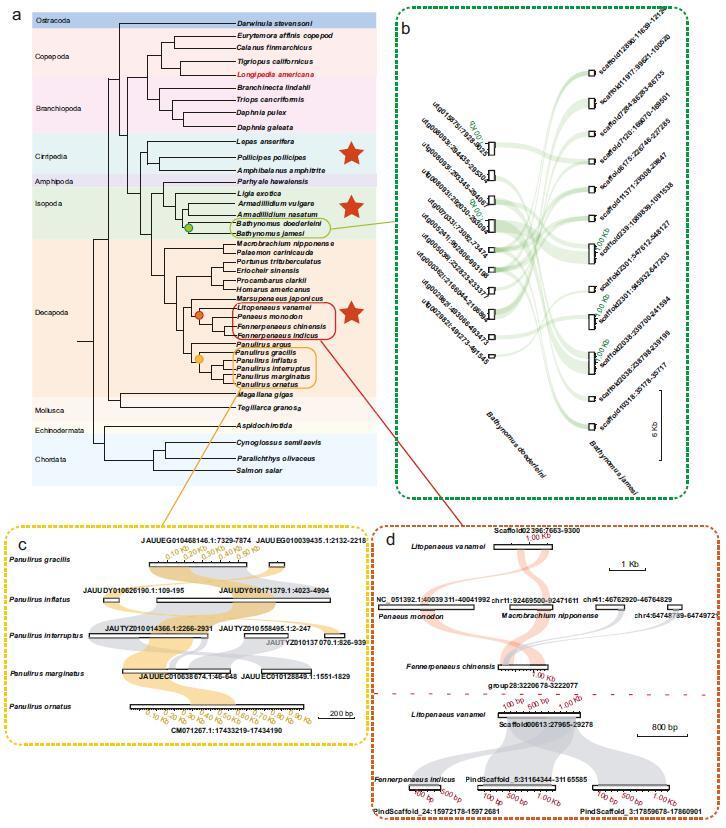
Fig. 1 Integration of IHHNV in the crustacean genome. (Image by IOCAS)
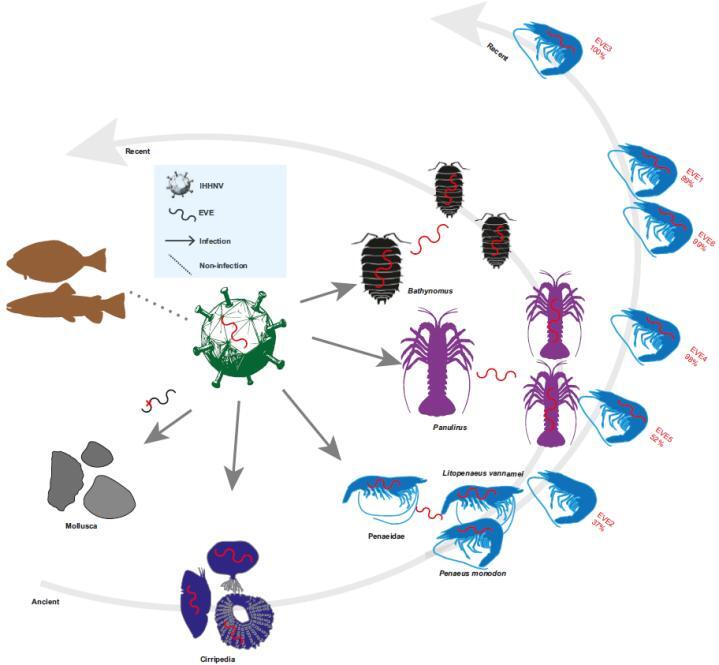
Fig. 2 Dynamic integration model of IHHNV-EVEs in Crustaceans. (Image by IOCAS)
(Text by LI Shihao)
Media Contact:
ZHANG Yiyi
Institute of Oceanology
E-mail: zhangyiyi@qdio.ac.cn
(Editor: ZHANG Yiyi)

