Linlin Zhang, Li Li, Ximing Guo, Gary W. Litman, Larry J. Dishaw & Guofan Zhang
Scientific Reports 5, Article number: 8693; doi:10.1038/srep08693
Received 24 September 2014; Accepted 02 February 2015; Published 03 March 2015
The molecules that mediate innate immunity are encodedby relatively few genes and exhibit broad specificity. Detailed annotation of the Pacific oyster (Crassostrea gigas) genome, a protostome invertebrate, reveals large-scale duplication and divergence of multigene families encoding molecules that effect innate immunity. Transcriptome analyses indicate dynamic and orchestrated specific expression of numerous innate immune genes in response to experimental challenge with pathogens, including bacteria, and a pathogenic virus. Variable expression of individual members of the multigene families encoding these genes also occurs during different types of abiotic stress (environmentally-equivalent conditions of temperature, salinity and desiccation). Multiple families of immune genes are responsive in concert to certain biotic and abiotic challenges. Individual members of expanded families of immune genes are differentially expressed under both biotic challenge and abiotic stress conditions. Members of the same families of innate immune molecules also are transcribed in developmental stage- and tissue-specific manners. An integrated, highly complex innate immune system that exhibits remarkable discriminatory properties and responses to different pathogens as well as environmental stress has arisen through the adaptive recruitment of tandem duplicated genes. The co-adaptive evolution of stress and innate immune responses appears to have an ancient origin in phylogeny.
The distribution in phylogeny of the cells, molecules and interactive processes that effect immune protection is of broad biological interest. In vertebrates, innate immunity, as well as lymphocyte-mediated adaptive immunity, is mediated by various classes of molecules and function in different phases of the response to foreign challenges. Adaptive immunity is a shared character of all vertebrates and is mediated through the somatic rearrangement of genes that give rise to genetically unique receptors expressed on the surface of individual lymphocytes1, 2. Innate immunity is a shared character of both vertebrates and invertebrates and relies on recognition of conserved pathogen-associated molecular patterns (PAMPs) present in microbes by germline encoded pathogen-associated pattern recognition receptors (PAMRs) including: toll-like receptors (TLR), retinoic acid-inducible gene I [RIG-I]-like receptors (RLRs) and NACHT-leucine-rich repeat receptor (NLR)3. Upon PAMP recognition, PAMRs activate intracellular signaling pathways, including adaptor molecules, kinases, and transcription factors and trigger proinflammatory and antimicrobial effectors4.
A number of different genetic mechanisms increase the diversity and specificity of the innate response of invertebrates, including: massive alternative splicing of Down syndrome cell adhesion molecule in fruit fly (Drosophila melanogaster)5, hypervariation and somatic variation in the fibrinogen-related proteins of the snail (Biomphalaria glabrata)6 and high allelic diversity in the immunoglobulin (Ig) domains of amphioxus (Branchiostoma floridae) variable region-containing chitin-binding proteins7 increase the diversity and specificity of the innate response of invertebrates. In both sea urchin (Strongylocentrotus purpuratus) and amphioxus8, 9, the diversity and likely specificity of immunity is achieved through large-scale expansion and diversification of multigene families encoding innate immune genes. The mechanisms underlying the expansion and functional diversification of the molecules at the transcriptional level are not understood.
Stress conditions, including challenges to the immune system, also are rapidly changing and highly variable. Stress adaptation likely requires rapid adaptive innovation10. Certain immune genes have been shown to have significant roles during abiotic stress responses, including: social stress in primates11, nutritional stress in the fruit fly12 and temperature stress in the alfalfa leafcutting bee13. Functionally divergent immune genes that are expressed during abiotic stress and/or biotic defense may share common pathways and are of broad general interest.
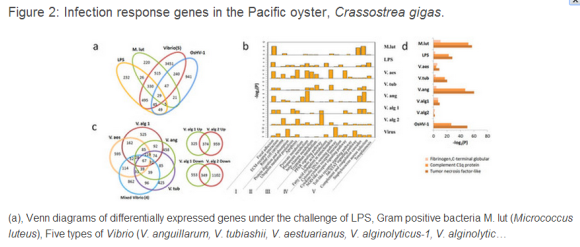
The Pacific oyster (Crassostrea gigas) is a member of the lophotrochozoa, a group of protostomes representing a large taxonomic group encompassing several major invertebrate taxa such as the Mollusca. As a sessile, filter-feeder exposed to a wide range of biotic (bacterial and viral) and abiotic stresses (dynamic variation in temperature, salinity and prolonged desiccation), the oyster represents an attractive model for studying the relationship of immunity and stress adaptation14and complements findings from an important lophotrocozoan system, B. glabrata, from which it was recently shown that a large proportion of transcripts are challenge-specific with some level of functional divergence noted from expanded gene families15. We present here a comprehensive genomic annotation and transcriptomic analyses of the large subset of genes that constitute the oyster immune system and determine that the lineage-specific expansion of genes is associated not only with differential responses to pathogens but also with differential expression under environmental stress conditions that emulate those of its natural habitat, as well as during the course of developmental maturation.
http://www.nature.com/srep/2015/150303/srep08693/full/srep08693.html
|
|

Address: 7 Nanhai Road, Qingdao, Shandong 266071, China
Tel: 86-532-82898902 Fax: 86-532-82898612 E-mail: iocas@qdio.ac.cn


