Sea cucumbers form one class of the echinoderms, which are all marine animals. Together with chordates and hemichordates, they form the deuterostome clade, making them a crucial node in the study of chordate ancestry. Besides, echinoderms also play important role in marine ecology and socioeconomic status. Among them, the sea cucumber is one of the most studied echinoderms due to its potent nutritional and medicinal properties. In China alone, around 200,000 tons of sea cucumbers were produced in the last few years.
To explore the genetic underpinnings of the sea cucumbers, the authors performed high-definition genomic sequencing of the sea cucumber Apostichopus japonicus, covering about 805 megabases of DNA, with the assembly scaffold N50 of 486 Kb. By comparing the genome of A. japonicus with that of other organisms, the authors found evidence that the echinoderms diverged from hemichordates about 533 million years ago and the sea cucumbers split off from other the echinoderm classes about 479 million years ago. They identified 763 echinoderm-specific gene families enriched in genes encoding membrane proteins, ion channels, and signal transduction proteins. Marker genes associated with the notochord and gill slits were also found, providing valuable insight into the origin of chordates. Sea cucumbers are unique among echinoderms in not having a hardened calcium exoskeleton. The authors showed that while the sea urchin genome includes 31 genes for biomineralization, critical for forming a calcified skeleton, the sea cucumber has only seven such genes. They also found that the sea cucumber expressed these biomineralization genes at much lower levels throughout development, likely accounting for their softer bodies compared to sea urchins.
As a strategy to scare off predators, sea cucumbers can expel their viscera, which they can then regenerate within several weeks. Genome, transcriptome, and proteome analyses of organ regrowth after induced evisceration provided insight into the molecular underpinnings of visceral regeneration, including a specific tandem-duplicated prostatic secretory protein of 94 amino acids (PSP94)-like gene family and a significantly expanded fibrinogen-related protein (FREP) gene family.
This high-quality genome resource will provide a useful framework for future research into biological processes and evolution in deuterostomes, including remarkable regenerative abilities that could have medical applications. Moreover, the multiomics data will be of prime value for commercial sea cucumber breeding programs.
This work was funded by the National High-Technology Research and Development Program of China, Strategic Priority Research Program of the Chinese Academy of Sciences, National Natural Science Foundation of China, and the Scientific and Technological Innovation Project of Qingdao National Laboratory for Marine Science and Technology.
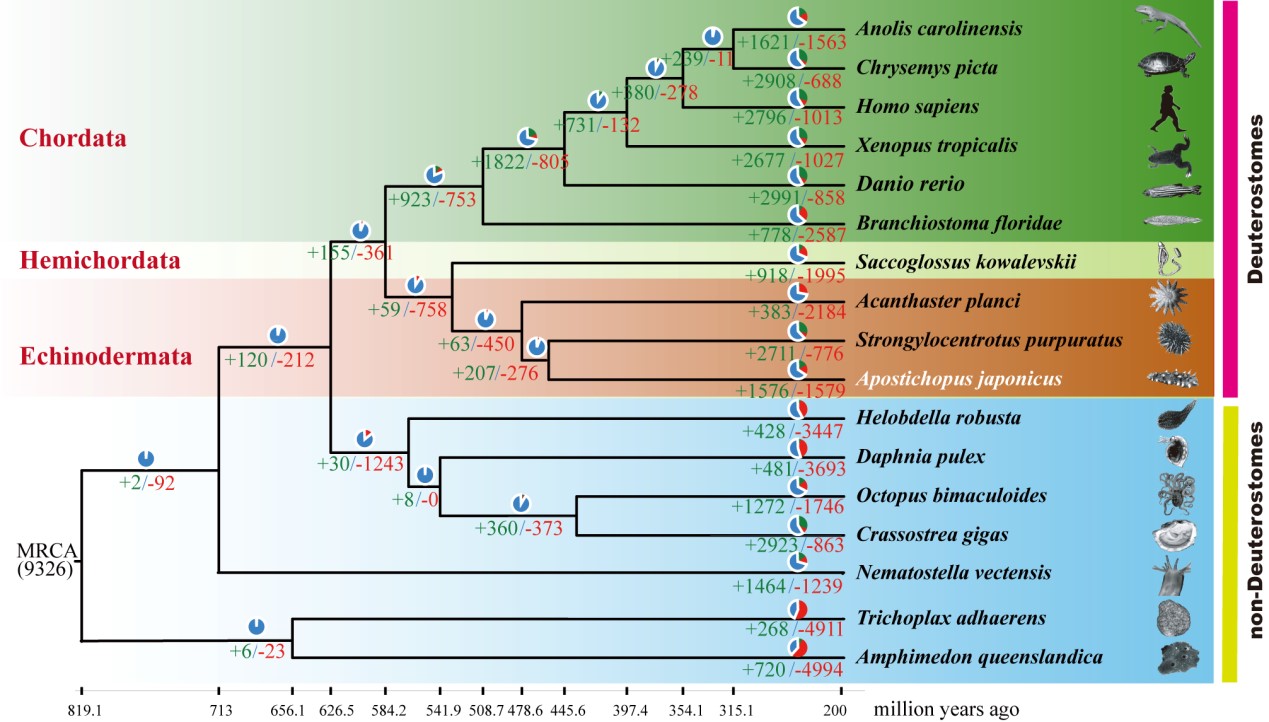
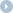
Fig 1. Phylogenetic tree of sea cucumber
Available article in PLOS Biology:
http://journals.plos.org/plosbiology/article/authors?id=10.1371/journal.pbio.2003790
|
|

Address: 7 Nanhai Road, Qingdao, Shandong 266071, China
Tel: 86-532-82898902 Fax: 86-532-82898612 E-mail: iocas@qdio.ac.cn


